A prescription for safer drugs
-
- from Shaastra :: vol 01 edition 01 :: May - Jun 2021
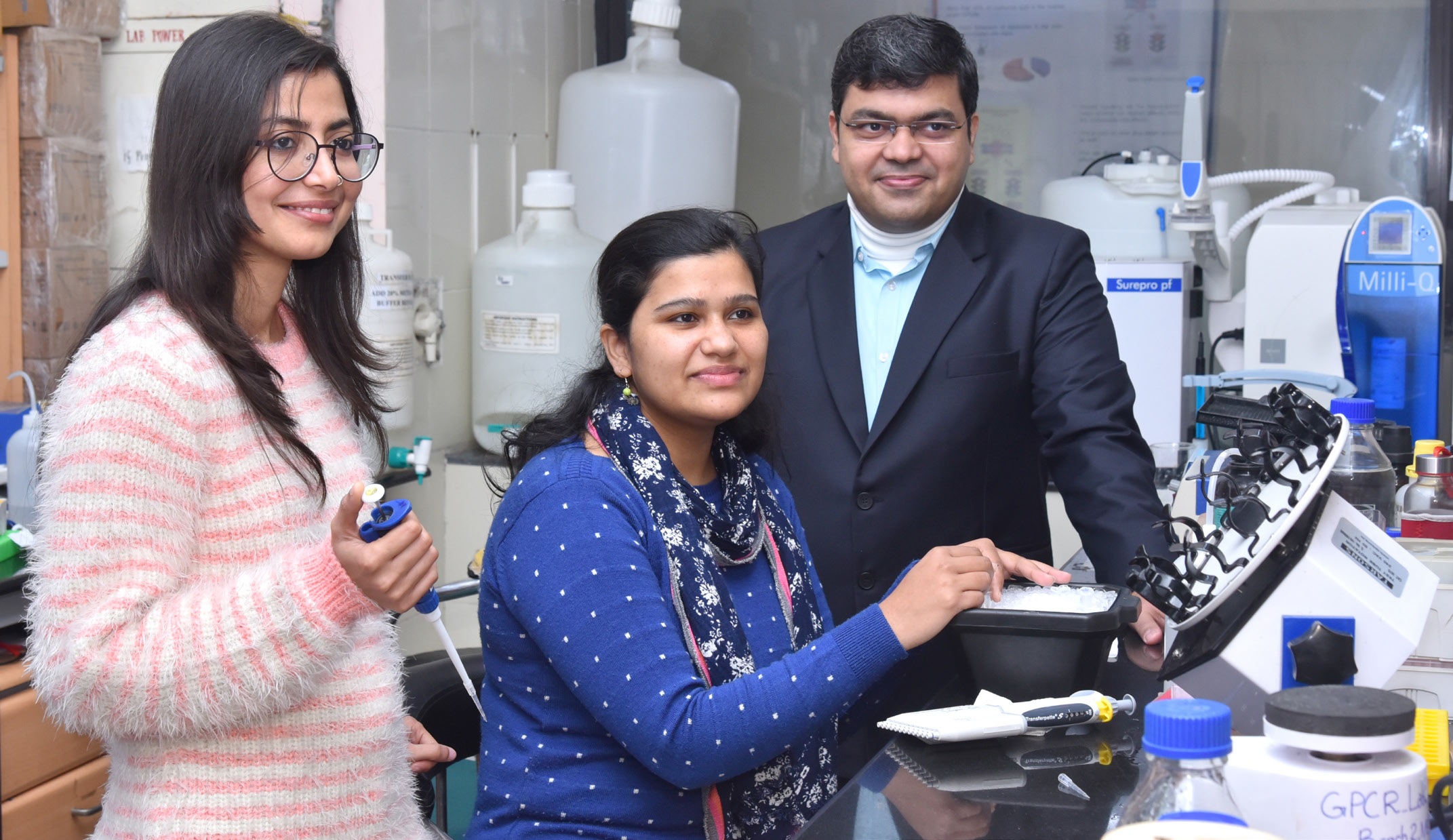

A path-breaking study, led by scientists at IIT Kanpur, uncovers a critical protein mechanism that holds promise for regulating drug effects.
Developing drugs that don’t have adverse side effects is every chemist’s dream. In August 2020, a small beginning was made towards this lofty goal when the US Food and Drugs Administration approved an opioid painkiller that belonged to a new class of drugs that can selectively activate only one subset of receptors called G protein-coupled Receptors (GPCRs). This drug, Oliceridine, developed by US-based biopharma firm Trevena, evidently works by preferentially stimulating signalling pathways responsible for therapeutic effect, while reducing – if not cutting off – the activation of those associated with adverse effects.
Nearly half the drugs that are in use today work through GPCRs. But there is a problem. These drugs activate not only GPCRs, but also an associated signalling pathway, called beta-arrestin 2, which is often linked to adverse effects associated with medication. For instance, with traditional opioid drugs, from which the Trevena drug Oliceridine is derived, these adverse events are related to respiratory and gastrointestinal complications.
The development of the Trevena drug owes much to Arun K. Shukla, a structural biologist at the Indian Institute of Technology (IIT) Kanpur, even though he did not play a direct role in it. That’s because Shukla has been among the few scientists who contributed immensely to the fundamental understanding of how GPCR and beta-arrestin 2 pathways work in tandem when a drug molecule is presented to a cell surface. His subsequent work has been instrumental in unravelling how the beta-arrestin triggers unwanted side effects.
Present on the surface of each cell in the human body, GPCRs act as “guardians of the senses”. GPCRs are as diverse as their functions are. They regulate everything from our sensory perceptions – the whiff of perfume, the unique taste of umami – to our heartbeats, the working of our brain, our ability to fight infections, and even the cheeriness of our spirit.
Much of the pioneering research work in the study of GPCRs had been carried out by biochemist Robert J. Lefkowitz and his collaborator Brian Kobilka, both of whom received the Chemistry Nobel in 2012.
Shukla’s interest in this area had drawn him, as a post-doctoral researcher in 2006, to Lefkowitz’s lab at Duke University, and to collaborate with Kobilka’s laboratory at Stanford University. Currently an Associate Professor and Joy Gill Chair at IIT Kanpur, Shukla has built on the illustrious legacy of his exceptional mentors, and his lab at the Department of Biological Sciences and Bioengineering (BSBE) has contributed to research work of enormous significance in this area.
Given the centrality of GPCRs to virtually every aspect of human physiology, they have become pivotal to the pharmaceutical industry. And it is this area that the most recent research work from Shukla’s lab, the findings of which were published in Science Advances journal in September 2020, addresses.
The study uncovers structural footprints in GPCRs that may be utilised to design safe drugs. It describes special sites (amino acids) in the human vasopressin receptor, a kind of GPCR, which are critical to modulating a mechanism that is central to regulating GPCRs-mediated drug effects.
COMPLEX BIOCHEMISTRY
The underlying science of this is a little complex. GPCRs are activated after binding to ligands or agonists (molecules that bind to other proteins) to send signals inside a cell. GPCR signalling occurs through the agency of two main actors: G-proteins, which bind activated GPCRs to initiate processes such as action of drugs; and beta-arrestin (βaar), which acts as a terminator of GPCR signalling and returns the receptor to its original deactivated state after its job is done. G protein-coupled receptor kinases (GRKs) act as supporting actors to add phosphate molecules to GPCRs (by way of a process called phosphorylation). βaar binds to the phosphorylated GPCR and stops the signalling.
Curiously, βaar can take one of two routes after binding to GPCRs. It can either stop the GPCR activity, or initiate a separate signal to yield a different biological outcome. To put this in the context of drug action, it bears considering that there are about 1,000 GPCRs, many agonists and multiple drugs. GPCR activation upon binding a drug may kick in signalling through both the G-protein arm and an independent βaar arm. This may result in biological responses that may yield beneficial effects or side-effects, which adds to the element of complexity.
The smart workaround would be to alter the receptor or drug selectively such that only the beneficial arm is activated. This is the concept of ‘biased signalling’, wherein a synthetic drug is created. These understandings of GPCR-βaar interactions made Shukla and his fellow researchers conjecture that cracking the arrestin code would be the key.
To do that, the goal is to visualise the GPCR-βaar complex at atomic resolution. Shukla explains: “Using cryo-electron microscopy, you will be able to see which parts of the receptor bind βaar. One starts with purifying the receptor and βaar, allows the two to form a stable complex in a test tube and image the structure using cryo EM.” Mithu Baidya, a post-doctoral fellow at Shukla’s lab, who was part of the latest study, says: “This is seminal work; the study describes the structural basis of GPCR-induced βaar activation.”
THE PROTEIN STRUCTURE
To understand the binding process, it helps to know the structure of the protein. Every protein is made up of chains of amino acids, with C-terminal and N-terminal ends, and a core domain. The layers, sheets and folds of amino acids give the protein its unique structure and function. Understanding the active structure allows one to selectively locate which amino acids or parts of the protein interact or bind with other proteins, and which changes in structure facilitate the binding or dissociation of a protein to/from another one.
A few amino acids, named Serine (Ser) and Threonine (Thr), are altered through a process of phosphorylation. This modification may govern how GPCR activates G-proteins and βaars by forming stable complexes. βaars bind to different phosphorylated amino acids in GPCRs to achieve multiple conformations, which may lead to varied functional outcomes. If one can map out these sites, the function of βaars may be modulated to the advantage of biasing the GPCR signalling.
Since 2013, when Shukla joined IIT Kanpur as a research group leader, his team has carried out several research studies in this space. In the first of them, researchers Punita Kumari and others demonstrated that binding of the N-terminal region of βaar to the C-terminal region of GPCR is enough to form a stable βaar-GPCR complex to carry out βaar-mediated downstream functions (Nature Communications, 2016).
That study broke new ground for membrane proteins structural biology research in India. Shukla then put together a team and mentored it for functional characterisation of phosphorylated amino acids in the C-terminal of GPCRs. How does one work out the role for a single phosphorylation site or a group of amino acids?
The best way is to change the amino acid(s) in GPCRs and see what happens to βaar activation. This could distinguish the sites that may lead to G-protein-based signal from the sites that allow an independent signalling via βaar. Researchers Hemlata D. Agnihotri and Madhu Chaturvedi led the task of bombarding the C-terminal to create mutations. Shubhi Pandey, a Ph.D. student, recalls: “We mutated many phosphorylation sites in the GPCR C-terminal.”
Baidya adds: “We first mutated a cluster of amino acids TSS (Thr369-Ser370-Ser371), but did not notice any effect on βaar activation and binding. However, when we touched another cluster SSS (Ser362-Ser363-Ser364), we saw a totally disrupted βaar activation. It was akin to carpet bombing.”
THE IMPLICATIONS
This study has established a paradigm on the unique role of phosphorylation sites in GPCR-induced βaar activation at the resolution and specificity of a single site.
Anurag Agrawal, asthma researcher and Director, CSIR-Institute of Genomics and Integrative Biology, says: “Their work helps us better understand the framework of GPCR-βaar interaction and biased signalling and opens up the path towards more effective therapeutics.” Shukla and his team have mapped the vasopressin receptor as a prototype. The same concept can be applied to any GPCR.
A group of researchers at Stanford University has published similar results (Cell, December 2020) using simulation approaches to demonstrate how phosphorylation sites in GPCRs may affect βaar activation, binding and signalling. Baidya says with justifiable pride: “Our wet lab results align with the dry lab simulation approaches.”
The ultimate goal of such research is to develop drugs. “We need to know how the receptor looks and how it works and then design the drug to test in animals,” says Sai Prasad Pydi, Assistant Professor at the Department of BSBE at IIT Kanpur. “Starting from the bench to bed, we need this whole spectrum of work. (Shukla’s) research takes care of the first part,” he adds. With the US drug regulator approving the first biased ligand-μ-opioid agonist oliceridine for pain management in adults, GPCR ligand biasing could definitely shape the future.
At IIT Kanpur, Shukla will soon have a Cryo EM facility funded by the Science and Engineering Research Board. He plans to study multiple drug-GPCR-arrestin complexes and unravel new structural masterpieces to realise his long-term vision of designing better drugs.
Have a
story idea?
Tell us.
Do you have a recent research paper or an idea for a science/technology-themed article that you'd like to tell us about?
GET IN TOUCH